 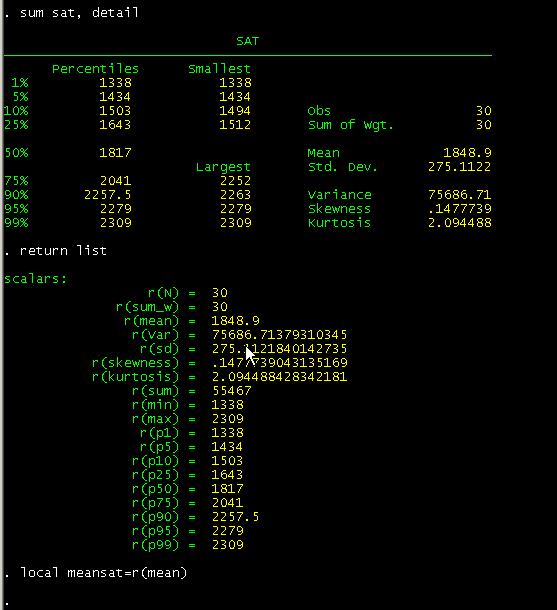
*(c) Plot the raw outcome data and the fitted model over time Glm numdeaths i.elapsedmonth, family(poisson) scale(x2) eform *(b) Fit elapsed month into the model with indicator variables ("i.") Gen elapsedmonths =(year -1996) *12 +month *(a) Generate a variable containing "elapsed month" (i.e. *Option 1 - Time-stratified model (simple indicator variables) *Modelling seasonality and long-term trend in the outcome data (leaving out the main exposure for now) Label var ozone10 "Ozone level in ug/m3 divided by 10 " 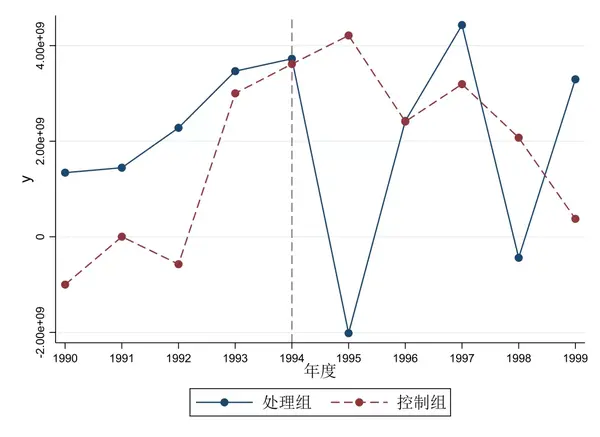
*Divide the ozone variable by 10 so that model estimates refer to a "per 10ug/m3 increase" (as per convention) Pwcorr ozone temperature relative_humidity *Correlation matrix for the explanatory variables Summ ozone temperature relative_humidity, d Graph combine deaths ozone, cols(1) xcommon name(Fig1, replace) Graph twoway scatter numdeaths date, msize(vsmall) mcol(black) name(deaths, replace) ytitle(Daily number of deaths) xtitle(Date) xline(`datelines', lp(dash) lc(gs10)) title(Daily deaths over time) Graph twoway scatter ozone date, msize(vsmall) mcol(gs5) name(ozone, replace) ytitle( "Daily mean ozone level (ug/m3) ") xtitle(Date) xline(`datelines', lp(dash) lc(gs10)) title(Ozone levels over time) *(b) Plot the exposure and outcome data against time, and combine the plots *(a) Create a list of dates to add as dashed guidelines in the plots 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
